- Who We Are
- Updates & News
- Standards
- Software Tools
- Network Studies
- Community Forums
- Education
- New To OHDSI?
- Community Calls
- Past Events
- Workgroups
- Tutorials
- 2025 ‘Our Journey’ Annual Report
- Current Events
- Support & Sponsorship
- 2025 Global Symposium
- 2026 Europe Symposium
- 2026 Global Symposium
- Github
- YouTube
- X/Twitter
- Newsletters
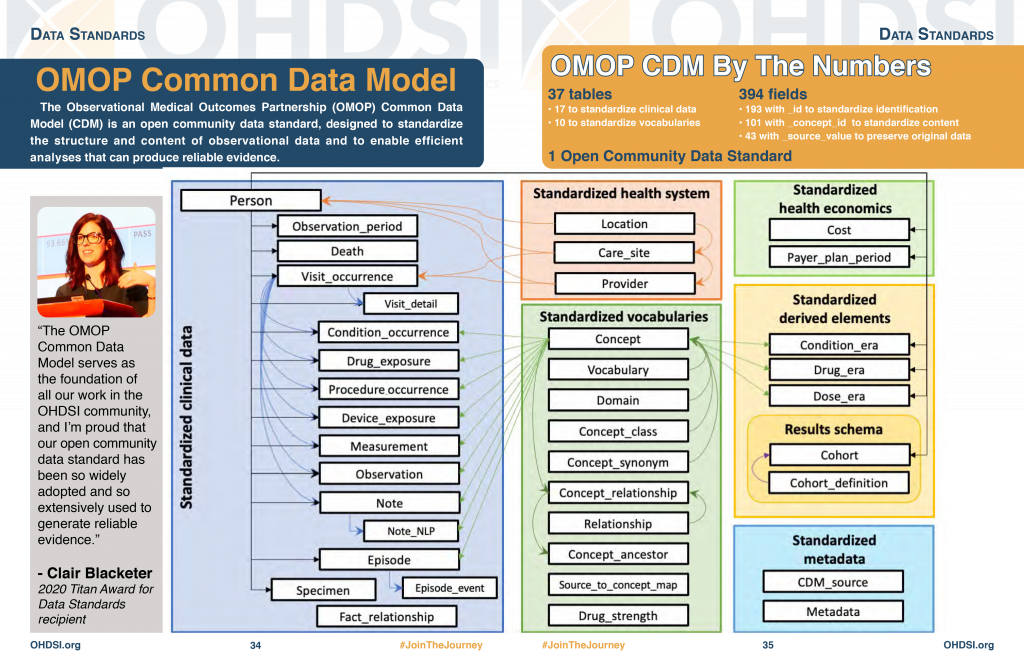
Standardized Data: The OMOP Common Data Model
The Observational Medical Outcomes Partnership (OMOP) Common Data Model (CDM) is an open community data standard, designed to standardize the structure and content of observational data and to enable efficient analyses that can produce reliable evidence. A central component of the OMOP CDM is the OHDSI standardized vocabularies. The OHDSI vocabularies allow organization and standardization of medical terms to be used across the various clinical domains of the OMOP common data model and enable standardized analytics that leverage the knowledge base when constructing exposure and outcome phenotypes and other features within characterization, population-level effect estimation, and patient-level prediction studies.
Data standardization is the critical process of bringing data into a common format that allows for collaborative research, large-scale analytics, and sharing of sophisticated tools and methodologies. Why is it so important?
Healthcare data can vary greatly from one organization to the next. Data are collected for different purposes, such as provider reimbursement, clinical research, and direct patient care. These data may be stored in different formats using different database systems and information models. And despite the growing use of standard terminologies in healthcare, the same concept (e.g., blood glucose) may be represented in a variety of ways from one setting to the next.
We at OHDSI are deeply involved in the evolution and adoption of a Common Data Model known as the OMOP Common Data Model. We provide resources to convert a wide variety of datasets into the CDM, as well as a plethora of tools to take advantage of your data once it is in CDM format.
Most importantly, we have an active community that has done many data conversions (often called ETLs) with members who are eager to help you with your CDM conversion and maintenance.
What is the OMOP Common Data Model (CDM)?
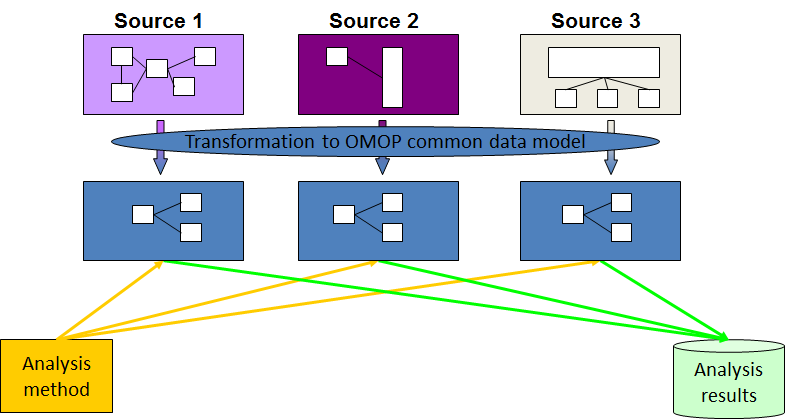
The OMOP Common Data Model allows for the systematic analysis of disparate observational databases. The concept behind this approach is to transform data contained within those databases into a common format (data model) as well as a common representation (terminologies, vocabularies, coding schemes), and then perform systematic analyses using a library of standard analytic routines that have been written based on the common format.
Why do we need a CDM?
Observational databases differ in both purpose and design. Electronic Medical Records (EMR) are aimed at supporting clinical practice at the point of care, while administrative claims data are built for the insurance reimbursement processes. Each has been collected for a different purpose, resulting in different logical organizations and physical formats, and the terminologies used to describe the medicinal products and clinical conditions vary from source to source.
The CDM can accommodate both administrative claims and EHR, allowing users to generate evidence from a wide variety of sources. It would also support collaborative research across data sources both within and outside the United States, in addition to being manageable for data owners and useful for data users.
Why use the OMOP CDM?
Once a database has been converted to the OMOP CDM, evidence can be generated using standardized analytics tools. We at OHDSI are currently developing Open Source tools for data quality and characterization, medical product safety surveillance, comparative effectiveness, quality of care, and patient-level predictive modeling, but there are also other sources of such tools, some of them commercial.